
SATURNA
AI for RNA BIOLOGY
Deciphering RNA structure, modification and function for therapeutic development
SATURNA is a research initiative dedicated to decoding the dynamic choreography of RNA, and to understanding how the process itself shapes disease states at cellular and molecular scales.
By combining AI innovation with RNA-focused, context-aware biology, SATURNA seeks to establish actionable principles of disease mechanisms, enabling new strategies to diagnose and treat complex human disorders.
SATURNA brings together expertise in artificial intelligence, computational RNA biology, molecular, cellular and clinical neuroscience. Within this framework, AI models are co-developed with domain experts to unlock concrete biological questions and accelerate their translation to clinic.

MISSIONS
SATURNA is built around four core missions:
Advance RNA science for human health
We conduct fundamental and translational research on RNA dynamics to address urgent human disorders, bridging basic discovery with clinical relevance.
Develop open and accessible AI technologies
We create novel AI methods which we release openly to ensure broad access, reuse, and impact across the scientific community.
Train the next generation of interdisciplinary leaders
We nurture aspiring molecular and cellular biologists, clinicians, and computer scientists in interdisciplinary research at the interface of AI and RNA biology. Our training program includes a network of PhD students and postdoctoral researchers, often co-supervised across disciplines and institutions.
Build community and engage society
We foster an inclusive international community and actively engage with clinicians, academics, industry partners, regulators, and the public. Through workshops, symposia, and public dialogue, we promote collaborative research as a key driver of responsible and impactful innovation.

FRAMEWORK
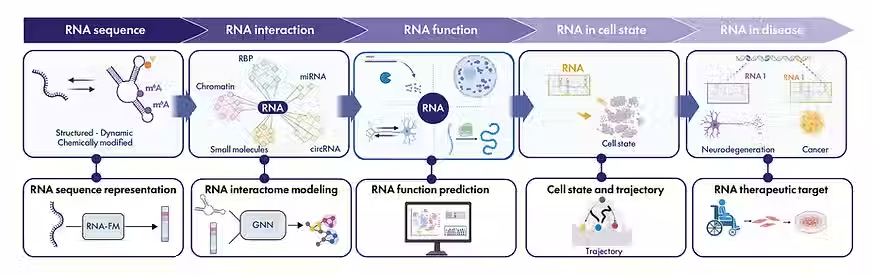
SATURNA’s scientific program is organised as a two-dimensional matrix integrating enabling technologies with three scientific pillars, with the goal of decoding RNA in context and delivering actionable RNA-driven therapies for complex human diseases, including cancer and neurodegeneration.

SCIENTIFIC OBJECTIVES
Advancing AI for RNA Biology
Building intelligent models to unravel the complexity and dynamics of RNA biology.
Advance state-of-the-art AI algorithms that integrate multimodal biological data, spanning molecular, cellular, imaging, and clinical domains with the vast amount of publicly available data and biological knowledge data-bases to build predictive and generative models of RNA sequences in context.
Decoding RNA function and regulation in a context-specific manner
Revealing how RNA is regulated and functions across cellular compartments, states and disease progression.
By combining advanced AI technologies with expertise in computational RNA biology, we explore emerging paradigms, from the influence of non-coding RNA elements on chromatin architecture to their role in protein compartmentalization, and their roles in neurodegeneration, cancer progression, and therapy resistance.
Engineering RNA-based therapeutics
Translating discovery into impact, from target identification to functional validation.
Non-coding regions of mRNA play critical roles in transcriptional diversity and disease, yet remain largely untapped as therapeutic targets. Leveraging our expertise in RNA biology and drug development, we aim to identify pathogenic RNA sequences underlying cancer and neurodegeneration and engineer precision therapeutics designed to address the earliest molecular and cellular drivers of disease.
SCIENTIFIC PILLARS
Language Models for RNA sequences
Translating large language models to learn biologically meaningful representations of RNA sequences and decode the underlying “grammar” of RNA biology.
Scientific Leader: Prof. Lonneke van der Plas — Università della Svizzera italiana, Lugano, Switzerland.
We leverage advanced AI methods, including but not limited to large language models (LLMs), to learn informative representations of RNA sequences. By modeling RNA as a structured language, we seek to uncover the deep features and contextual dependencies embedded within its sequence. In parallel, we interpret what these models learn, probing the internal representations to understand what biological information they capture, how they encode structure, regulation, and function, and what this reveals about the underlying grammar of RNA biology.
Graph-based machine learning and signal processing
Modeling RNA interactomes and disease trajectories using graph and generative models across multimodal cellular data
Scientific Leader: Prof. Pascal Frossard — EPFL, Lausanne, Switzerland.
We develop self-supervised and generative graph representation learning to model context-specific, multimodal RNA interactomes and develop robust modeling of disease trajectories. By modeling RNAs within their relational and functional contexts, we aim to improve the representation of RNA sequences and enable reasoning over complex biological networks. This approach allows us to explore how RNAs interact with other molecular entities, such as proteins, metabolites, or other RNAs, across diverse cellular environments, thereby advancing our understanding of RNA functions in context and their roles in health and disease. We also develop conditional generative modeling of RNA sequences with defined regulatory properties to advance RNA-based therapeutics.
Knowledge-Augmented RNA modeling
Integrating ontologies and knowledge graphs into AI systems to enhance interpretability, generalization, and context-aware RNA predictions.
Scientific Leader: Dr Janna Hastings — Idiap Research Institute, Martigny, Switzerland.
Injection of biological knowledge into AI technologies has the potential to significantly improve performance and generalisability of RNA-LMs by incentivising models to learn the established biological principles underlying common patterns. We develop and evaluate a methodology to inject a wide range of prior curated knowledge about RNA structural motifs, functional domains and regional evolutionary conservation into RNA-LMs.
Computational RNA biology
Decoding how RNA structure, modifications, and interactions regulate cellular organization and disease-associated state transitions.
Scientific Leader: Prof. Raphaëlle Luisier — University of Bern, Bern, Switzerland.
We leverage existing data and tools developed in this program to study the functional roles and regulatory mechanisms of RNA. We are particularly interested in elucidating the function of non-coding mRNA regions in driving the subcellular compartmentalization of biomolecules underlying physiological and disease cell-state transitions. We seek to decode the combinatorial rules that govern RNA behavior, integrating information on RNA modifications, structural features, and molecular interactions. Using these learned representations and relational models, we investigate how RNAs contribute to subcellular compartmentalization and the organization of biomolecular condensates, providing insights into the spatial and functional logic of RNA regulation within the cell, eventually elucidating the function of non-coding mRNA regions in driving disease cell state transitions.
Experimental stem cell biology and RNA therapeutics
‘Uncovering new disease mechanisms, identifying and validating RNA-based biomarkers and therapeutic targets
Scientific Leader: Prof. Rickie Patani — National University of Singapore, Singapore.
We study the role of deregulated RNA metabolism in ALS, and identify therapeutic targets for ALS using collaborative insights and tools developed in this program. We generate spatially and temporarily resolved rich molecular data including CLIP-seq, RNA-seq, mass-spec and imaging data that strengthen the AI and computational methods developed in this program, reinforcing the synergies among out efforts.

LEADERSHIP
Founded by Prof. Raphaëlle Luisier, SATURNA brings together five leading laboratories with complementary expertise in artificial intelligence and RNA biology.
This international alliance was created to enable true interdisciplinary collaboration and accelerate the development of RNA-based therapies for urgent unmet medical needs, including neurodegeneration and cancer.


RESEARCHERS
Independent Researcher
Dr Cédric Vincent-Cuaz, Bern University, Bern, Switzerland
Postdoctoral researchers
Dr Charlotte Tumesheit, Idiap, Martigny, Switzerland
Dr Damien Leow Meng Kiat, NUS, Singapore
PhD students
Vincent Jung, EPFL, Lausanne Switzerland
Michael Jopiti, Unibern
Lisa Fournier, EPFL
Elisa Messori, EPFL
Interns
Alois Thomas, Bern University, Bern, Switzerland

COLLABORATORS
Recognising that RNA is ubiquitous and impacts virtually all diseases, we have developed a large network of national and international collaborators that complement our expertise and enable us to broaden our vision, ensure our tools can be leveraged to address additional disorders, and bring our research to the next level.
Pathology
Dr. Igor Letovanec, MD, Institut Central des Hôpitaux (ICH), Hôpital du Valais, Sion
Oncology
Prof. Intidhar Labidi-Galy, MD, PhD, Ovarian cancer, University of Geneva – HUG
Dr. Uli Simon Herrmann, MD, PhD, Wilms tumour, University of Bern – Inselspital
Prof. Mark A. Rubin, MD, Prostate cancer, University of Bern – Inselspital
Dr. med. et phil. nat. Ruben Bill, Breast cancer, University of Bern – Inselspital
Molecular biology
Prof. Antonella Riccio, RNA metabolism in neurons, MRC Laboratory for Molecular Cell Biology, UCL, London, UK
Prof. Marc-David Ruepp, RNA metabolism in neurodegeneration, UK DRI at King's College London, UK
Prof. Silvia Barabino, RNA biology & neurodegeneration, University of Milano-Bicocca, Italy
Prof. Pei-Hsuan Wu, RNA metabolism in fertility, Department of Genetic Medicine and Development, University of Geneva
Therapeutic Development
Dr Bernard Schneider, Viral vector & gene therapy development, Bertarelli Foundation Gene Therapy Core Facility, EPFL

PUBLICATIONS

Visions of the future of molecular cell biology. M Abu-Remaileh, CJ Chan, L Chen, GS Demirer, A Fiszbein, F Jug, A Victoria Lechuga-Vieco, R Luisier, J Pagan, B R Sabari, S Shao, L Sun, JJ Żylicz. Nature Reviews Molecular Cell Biology (2025)
To mark the journal’s 25th anniversary, 13 researchers envision how breakthroughs such as artificial intelligence-powered virtual cells could transform our understanding of biology and reshape therapies and sustainable agriculture by 2050. Link to the journal here.

Decoding the interactions and functions of non-coding RNA with artificial intelligence. V Jung, C Vincent-Cuaz, C Tumescheit, L Fournier, M Darsinou, ZM Xu, A Saadat, Y Wang, P Tsantoulis, O Michielin, J Fellay, R Patani, A Ramos, P Frossard, J Hastings, A Riccio, L Van der Plas, R Luisier. Nature Reviews Molecular Cell Biology (2025)
In this roadmap review, we propose a forward-looking framework that combines AI-driven model development with advanced molecular and clinical sciences to decode RNA within its biological context and accelerate the translation of RNA-based therapeutics into clinical impact. Link to the journal here.

Amyotrophic lateral sclerosis caused by TARDBP mutations: from genetics to TDP-43 proteinopathy. R Balendra, J Sreedharan, M Hallegger, R Luisier, HA Lashuel, JM Gregory, R Patani. The Lancet Neurology (2025)
This review explores the role of TDP-43 in amyotrophic lateral sclerosis (ALS), highlighting how its mislocalisation and aggregation drive neurodegeneration. It discusses underlying molecular mechanisms and emerging biomarker and therapeutic strategies aimed at enabling more personalised approaches to treatment. Link to the journal here.

Extended pre-training of histopathology foundation models uncovers co-existing breast cancer archetypes characterized by RNA splicing or TGF-β dysregulation. L Fournier, G Haefliger, A Vernhes, V Jung, I Letovanec, P Frossard, C Vincent-Cuaz, R Luisier. Preprint (2025).
This interdisciplinary study integrates artificial intelligence, computational pathology and molecular biology to specialize histopathology foundation models for invasive breast cancer. By linking tissue morphology with gene expression, it uncovers clinically relevant tumor archetypes and advances precision oncology using routine pathology images.
TOOLS
Pipeline to systematically evaluate the biological concepts encoded within histopathology foundation models (hFMs) using molecular data. We also perform extended-pretraining of UNI to identify optimal conditions that enhance the model’s ability to encode richer, tumor tissue-specific biological concepts.

FORUM
AI for Biomolecule symposium
The AI for Biomolecules Symposium is a SATURNA-led initiative co-chaired by Prof. Raphaëlle Luisier and Dr Janna Hastings. It brings together domain experts and computer scientists to discuss the latest advances in AI addressing key challenges in RNA and protein biology.
Each month, the symposium hosts a 2-hour hybrid mini-event featuring one international speaker and one early-career Swiss researcher (PhD student or postdoctoral fellow), fostering cross-disciplinary dialogue, visibility for young scientists, and sustained community building.

ART & SCIENCE
Discovery is not driven by data alone, it also requires imagination. We see art as a catalyst for new ways of thinking, helping us question assumptions, communicate beyond disciplinary boundaries, and embrace the complexity of the human experience.
Artists bring perspectives that science alone cannot capture. Through this dialogue, we open new channels of engagement with society while encouraging our researchers to cultivate creativity, broaden their outlook, and approach challenges with greater adaptability.
This exchange enables us to explore problems more deeply, share knowledge more openly, and co-create solutions that are not only scientifically rigorous, but also socially meaningful and profoundly human.
A bilingual work authored in four hands by Marousa Darsinou (English) and Raphaëlle Luisier (French).

JOIN US
SATURNA regularly hosts undergraduate and Master’s students (BSc, MSc) for 4–6 month internships and Master’s thesis projects. A current list of available projects can be found [here].
Interested students are invited to contact the relevant project lead or PI, including a brief description of their background and research interests.
We also welcome PhD and postdoctoral candidates. Applicants are encouraged to send their CV and a short statement of interest. SATURNA actively supports applications to competitive doctoral and postdoctoral fellowships and visiting scholars.
FUNDING
Our research is supported by:






Support Our Mission
SATURNA is driven by a belief that understanding RNA is key to unlocking new treatments for some of humanity's most complex diseases. We welcome partnerships with foundations, philanthropists, and industry partners who share this vision.
If you are interested in supporting our work — whether through research funding, collaborative projects, or philanthropic giving — we would love to hear from you.





